Başlık: Bartonella species in wild small mammals in Western Black Sea Region of Turkey Yazar(lar):ÇELEBI, Bekir; KARAGÖZ, Alper; ÖKTEM, Mehmet Ali; ÇARHAN, Ahmet; MATUR, Ferhat; ÖZKAZANÇ, Nuri Kaan; BABÜR, Cahit; KILIÇ, Selçuk; SÖZEN, Mustafa; KARATAŞ, A
Tam metin
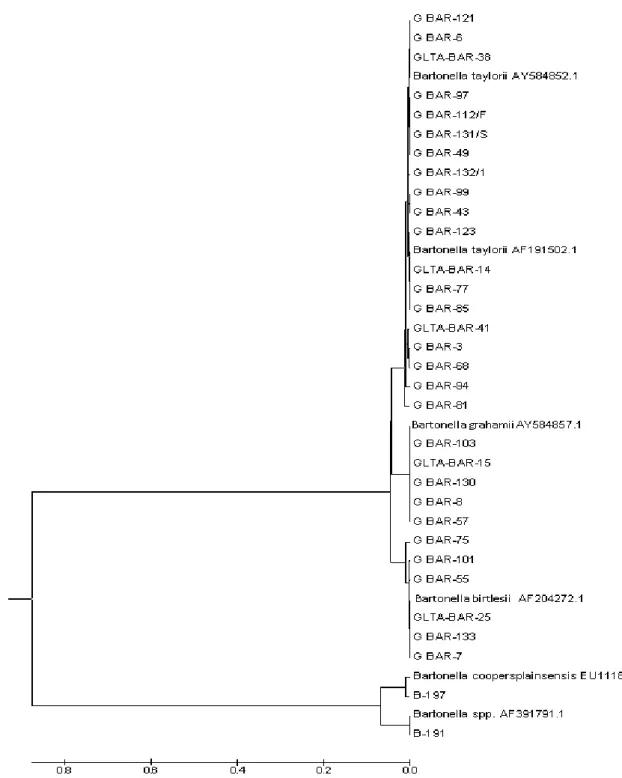
Şekil

Benzer Belgeler
This proceedings includes the 8 papers presented at the 6th IAPR Workshop on Pattern Recognition in Remote Sensing (PRRS 2010) that was held on August 22, 2010 in Istanbul, Turkey
The effects of the adsorbed carbon atoms C ∗ on the electronic structure of bare graphene are revealed by the calculations of energy band structure, total (TDOS) and projected
In an operating room with recirculating laminar flow and horizontal HEPA filter, optimal conditions have been investigated in order to obtain laminar flow form by
Araştırma arazisinin Çaylı serisi üzerinde yeni ve eski toprakaltı drenaj sistemlerinin yüzey toprağı infiltrasyonuna etkilerini araştırmak için yapılan
Dynamic contrast-enhanced Magnetic resonance imaging (DCE-MRI ) were classified Type 1: Progressive heterogeneous enhancement; Type 2: More pronounced enhancement towards
Yoğun bakım ünitelerinden bir yıl boyunca yapılan toplam istek sayısı ve transfüzyon merkezinden yapılan çıkıĢ sayısının kliniklere göre dağılımı incelendiğinde,
Zaferden sonra da Celâl Beyi yorulmuş görmedik, İzmir me busu iken, cumhuriyetin ilâ nından sonra en büyük yurt davalarından biri olan imar ve iskân işlerinin
Her bir tekrar boyunca elde edilen minimum kayıpların değişimleri Şekil 9’da, her bir tekrarın sonunda bulunan minimum kayıpların değişimleri Şekil 10’da,