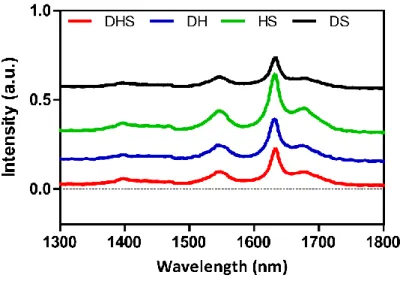
Catalytic and bioactive nanostructures for regenerative medicine applications
Tam metin
Şekil
Benzer Belgeler
Figure 3.7: A hierarchical drawing sample with compound nodes; note that graph is directed (courtesy of Tom Sawyer Software)....
As discussed in the Methodology section, obtained volumetric force maps simulating the interaction between tip and sample are “scanned” with a virtual NC-AFM based on the
There is no question today that norms of human rights pose a fundamental challenge to norms of state sovereignty especially in developing countries.. The debate between those who
That is, if the discount factor of the long-run player tends to 1 while holding the commitment type’s ex ante probability fixed, then the aforementioned reputation result à la
In this paper, we present a novel neural model and demonstrate that phase modulation can be achieved by inhibitory surround of feedback signals and it serves to reject
In our proposed design processes, the spinal (deepest) region of the scaffold architecture needs to be determined so that the gradient of functional porous structure can
also found that increasing the dosage of Pinus radiate (100-500mg) led to an increase in the % removal of methylene blue due from aqueous solution while a
To give information systematically and comprehensively about the enzymes which are biomolecules used for the reactions in biological systems and to show how
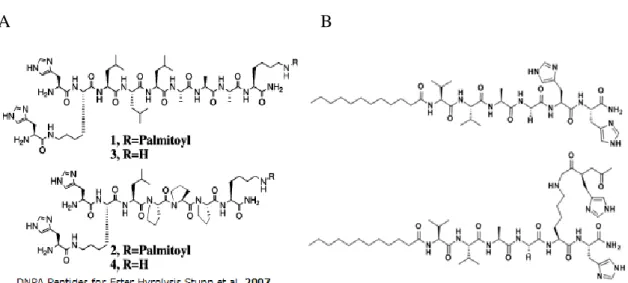
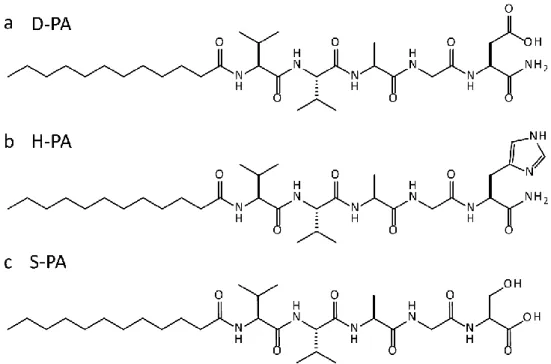
![Figure 2.2 Mass spectra of catalytic triad peptides. Following the subtraction of water readings, results found as a) For H-PA, [M+H] + (calculated) = 664.86, [M+H] + (observed) = 664.4967, [M/2+H] + (calculated) = 332.43, [M/2+H]+ (o](https://thumb-eu.123doks.com/thumbv2/9libnet/5623622.111447/67.892.169.834.128.660/catalytic-peptides-following-subtraction-readings-calculated-observed-calculated.webp)