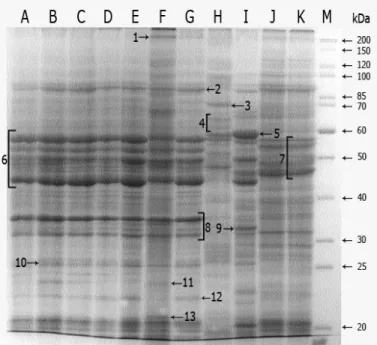
Discrimination and numerical analysis of human pathogenic Candida albicans strains based on SDS-PAGE protein profiles
Tam metin
Şekil
Benzer Belgeler
In this section, we present simulation results for the STBC- SM system with different numbers of transmit antennas and make comparisons with SM, V-BLAST, rate-3/4 OSTBC for
The findings from this pilot study suggest that the CTI-C, the SCS, and the CCLSS are valid and reliable instruments for measuring Adolescents' cognitive triad, self-control
Overall, macroeconomic announcements by both Bundesbank and the Federal Reserve are significant in explaining the volatility of the DEM/$ exchange rate for the 15-minute
[r]
Nestor İskender’in yazdığı kabul edilen “Çar Şehrinin Öyküsü (Şehrin Kuruluşu ve 1453 Yılında Türk- ler Tarafından Alınışı)” adlı tarih-savaş
• Proteinlerin transfer olduğu membran %5 BSA’lı TBST içinde (bloklama tamponu) 1.5 oda sıcaklığında uygun plastik kabın.. içerisinde
Mathematical examinations are performed utilizing a nonlinear limited component ( FE) analysis by joining damage plasticity model (CDP), for material conduct the
This also served as the arrival point for Dalit feminism, as ‘feminism’ in India — based as it was on the understanding of Brahmin/upper caste women due to their relatively